GenAI-Net (RL4CRN)
This repository contains the reference implementation of GenAI-Net — a generative AI framework for automated biomolecular / chemical reaction network (CRN) design using reinforcement learning and simulation-based evaluation. (arxiv.org)
Overview
GenAI-Net treats CRN design as a generative sequential decision process over a hybrid search space: - Discrete structure: which reactions (and thus which topology) the network contains - Continuous and discrete parameters: kinetic constants and other reaction-specific parameters - Optional input–species influence structure (when enabled)
A policy proposes edits to a candidate network, the resulting CRN is evaluated via deterministic and/or stochastic simulation against a task objective, and the agent learns to generate progressively better networks over time.
At a high level, GenAI-Net follows this loop:
- Observe the current CRN state (structure + parameters, optionally input influence)
- Act by proposing a CRN modification (e.g., add a reaction from a library + sample parameters)
- Simulate the modified CRN (ODE or SSA)
- Score it with a task-defined objective (reward / loss)
- Learn a policy that generates high-performing CRNs efficiently
GenAI-Net system overview
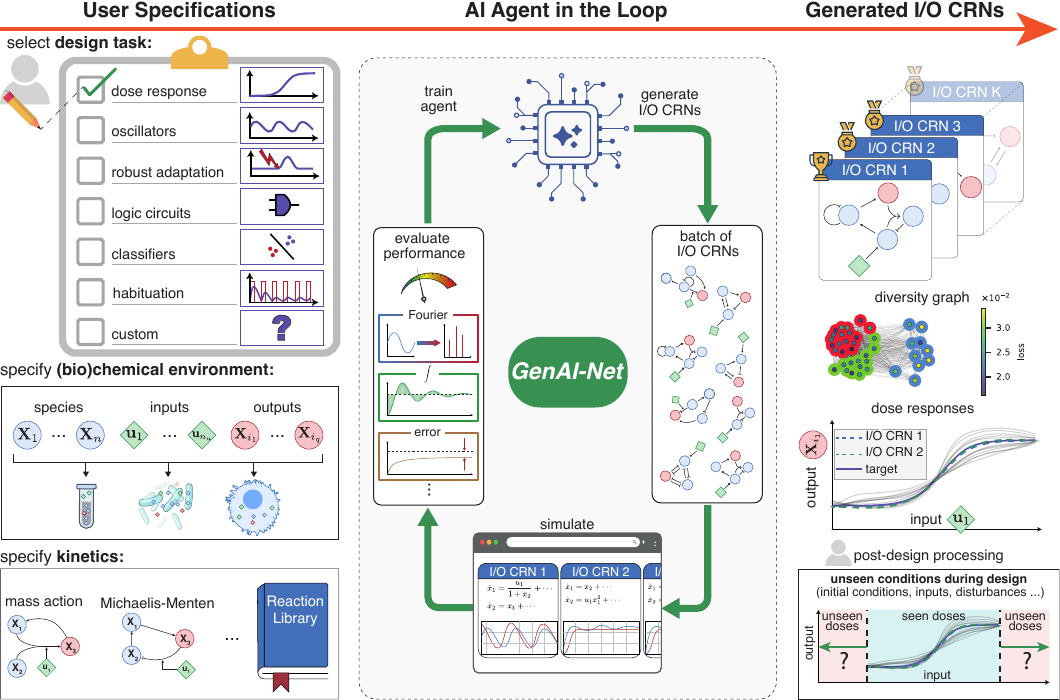
The end-to-end GenAI-Net pipeline. a policy generates CRN edits (topology + parameters), the candidate is evaluated in simulation under a user-defined task objective, and learning shifts the proposal distribution toward better-performing and diverse networks.
Code structure
The code follows the same conceptual decomposition as the GenAI-Net method:
-
iocrns/
Core CRN and IOCRN representations (species, reactions, reaction libraries, simulation hooks). -
env2agent_interface/
Observers and tensorizers: convert an environment/CRN state into the tensor representation consumed by neural policies. -
agent2env_interface/
Actuators and steppers: apply an agent action to mutate a CRN (e.g., add reaction, set parameters) and step the environment forward. -
policies/
Neural policies for proposing structure and sampling parameters (including distribution-backed parameter generators). -
distributions/
Distribution utilities used by policies (e.g., categorical/lognormal helpers, multivariate variants). -
agents/
RL algorithms that optimize policies using rewards returned by environments. -
environments/
Single and multi-environment wrappers, including serial and parallel execution. -
rewards/
Reward / loss functions for deterministic and stochastic simulations (tracking, oscillations, logic, robustness, etc.). -
utils/
Common utilities: FFNNs, initial-condition helpers, hall-of-fame storage, metrics, SSA summarization, and visualization tools.
Installation
Clone the repository and install in editable mode:
git clone <YOUR_REPO_URL>
cd <YOUR_REPO_DIR>
pip install -e .
Reference
Filo, M., Rossi, N., Fang, Z., & Khammash, M. (2026). GenAI-Net: A Generative AI Framework for Automated Biomolecular Network Design. arXiv preprint arXiv:2601.17582.